Tong Zhang, Environmental Biotechnology Laboratory, the University of Hongkong, China
Antibiotics have been discharged into wastewater treatment plants (WWTPs) for decades. As a result, multiple classes of antibiotics have been widely detected in different WWTPs worldwide. The widely applied biological treatment process creates favourable conditions for antibiotic resistance gene (ARG) development and horizontal gene transfer under the sub-inhibitory antibiotic concentrations. So far, six classes of antibiotics, i.e., β-lactams, sulfonamides, quinolones (fluoroquinolones), tetracyclines, macrolides and others have been detected in the influents and effluents from WWTPs worldwide. A big portion of ARGs in natural environments and fecal samples could be detected in WWTPs and the levels of ARGs in WWTPs are comparable to the fecal sources. These results demonstrated that the roles of WWTPs as hot spot for ARGs spread desire further comprehensive study to develop the integrated strategy to fight the resistance.
The dissemination of antibiotic resistance genes (ARGs) carried by antibiotic resistant bacteria (ARB) and their threats to public health have attracted concerns worldwide in the recent years (1). A number of organizations and governments have enhanced their monitoring schemes of ARB and ARGs, including the World Health Organization (2), the Public Health Agency of Canada, the Houses of Parliament of United Kingdom and the White House in the United States.
Besides those concerns on antibiotic resistance in clinical environments, interest has also arisen in ARGs entering the environment. Up until now, ARGs have been widely detected in various environments, including soil (3), water (4), sediments (5), where they have the potential to be transferred from host bacteria to pathogens by horizontal gene transfer (HGT). ARGs are now regarded as emerging pollutants and their dissemination in the environment has attracted wide attention. However, the source and fate of ARGs in the environment is still not fully understood yet. Wastewater treatment plants (WWTPs) are among the main anthropogenic sources of ARGs discharged into the environment. The widely applied biological treatment process, such as activated sludge, creates favourable conditions for ARGs development and HGT under the sub-inhibitory antibiotic concentrations.
Antibiotics as chemical pollutants in WWTPs
As chemical pollutants, antibiotics have been discharged into WWTPs for decades from different sources, including households (domestic), hospitals (clinical) and pharmaceutical factories (industrial). As a result, multiple classes of antibiotics have been widely detected in different WWTPs worldwide (6). So far, at least six classes of antibiotics, i.e., β-lactams, sulfonamides, fluoroquinolones, tetracyclines, macrolides and others have been detected in the influents and effluents from WWTPs (6). Concentrations for the same antibiotic in different sites may vary significantly, sometimes even by 1~2 orders of magnitude. Such significant variation of antibiotic concentrations in wastewater influent could be due to multiple reasons, including antibiotics consumption pattern, seasonal (even hourly) fluctuation, and the size of catchment area of the WWTP, etc. Antibiotic consumption patterns are quite different in different areas. Even in the same country, regional and local usage patterns may vary greatly (7), resulting in different concentrations of antibiotics in the WWTP. For example, cefotaxime was detected at low concentration (<15 ng/L) in sewage influents of Hong Kong while it was detected with a much higher mean concentration of 1100 ng/L in sewage influent of Shenzhen (China), a neighbouring city of Hong Kong. In contrast, cefalexin was lower than the detection limit in Shenzhen while its concentration was up to 1900 ng/L in sewage influent of Hong Kong. Secondly, the different sampling time may give very different results since the seasonal fluctuations of antibiotic consumption is high and also the hourly change in one day (8). Thus, different sampling procedures (grab/composite sampling, flow/time proportional sampling) will result in great concentration fluctuations. Lastly, the catchment area size may also affect the variation of antibiotic concentration. The less population the WWTP serves, the more the concentration fluctuates (8). Additionally, average daily mass flows indicated that the antibiotic consumption patterns and amounts varied significantly from region to region, even in the same city (8), depending on the populations in that catchment area.
Among β-lactams antibiotics, the highest concentrations detected in influent and effluent was up to 13800 ng/L (Penicillin V) and 2000 ng/L (Penicillin V) (9), respectively. Overall, although used in great amounts, β-lactams were not frequently detected (10), probably due to that β-lactam ring is unstable and can be cleaved by β-lactamases, a group of widespread enzymes in bacteria, or by chemical hydrolysis. Among sulfonamides, sulfamethoxazole was the most frequently detected, followed by sulfamethazine, sulfapyridine and sulfadiazine. The highest concentration of sulfamethoxazole was about 6000 ng/L in wastewater (11). The occurrence of quinolones in WWTPs was worldwide, consistent with its’ universal extensive usage. Twelve quinolones have been detected in WWTPs, including two first generation ones (pepemidic acid and nalidixic acid), eight second generation ones and two fourth generation ones (moxifloxacin and gatifloxacin) (6). Among these quinolones, ciprofloxacin and ofloxacin were the dominant ones with high detection frequency and high concentration, i.e., 4600 ng/L (ciprofloxacin) (12) and 7870 ng/L (ofloxacin) (13), respectively. Among macrolides, erythromycin-H2O had the highest concentrations, i.e., 10025 ng/L (14) and 4330 ng/L (13), in influent and effluent, respectively, followed by roxithromycin, clarithromycin, azithromycin and tylosin. Five tetracyclines have been detected in WWTPs with the highest concentration of 2210 ng/L (doxycycline) in influent (15) and 1420 ng/L (tetracycline) in effluent (13).
Although many individual surveys of antibiotics in wastewater treatment plants have been conducted, the systematic monitoring data of antibiotics and ARGs in typical municipal wastewater treatment plants are still limited. It is strongly suggested that surveys are conducted by taking frequent samples, including influent, supernatant of the primary sedimentation tank, mixed liquid in the aeration tank, supernatant in the secondary sedimentation tank, effluent after disinfection, rejected water from sludge dewatering, etc.
The major removal pathways of antibiotics in wastewater treatment processes include adsorption, biodegradation, disinfection as well as membrane separation. Other removal pathways, such as hydrolysis, photolysis and volatilization may be ruled out due to their minor role for antibiotics reduction in wastewater treatment processes. Some of the antibiotics (for example, β-lactam) could be eliminated by biodegradation while some of them (for example, tetracycline) would be adsorbed by activated sludge flocs or biofilm and largely concentrated there (16). For β-lactams, despite that it accounts for the highest proportion (50~70%) of the total human use antibiotic consumption, its occurrence was not detected frequently due to its unstable property (7). However, for those adsorbed by activated sludge or biofilm, although their concentrations in the bulk water of wastewater are relatively low (usually at the range of 0.1~1 µg /L), their concentrations in the activated sludge or biofilm could be as high as 0.1~1 mg/L, considering the high distribution coefficient (17). At this sub-inhibitory concentration, bacteria in activated sludge is not being killed or inhibited, but get the chance to develop resistance under the selective pressure (18).
To better understand the adsorption and biodegradation of antibiotics in activated sludge, some laboratory experiments have been conducted to investigate the adsorption and biodegradation (16). The results demonstrated that biodegradation and adsorption were the major removal routes for the antibiotics in activated sludge process under both aerobic and anoxic conditions. However, there is still very limited knowledge about the biodegradation and adsorption behaviours for most of the commonly used antibiotics.
Antibiotics resistance genes as biological pollutants in WWTPs
Activated sludge has been widely used as a biological wastewater treatment process for over 100 years and plays an important role in control of conventional pollutants, including suspended solid, BOD/COD, nutrients (N/P), etc. The bacterial diversity in activated sludge is very high. At the same sequencing depth (say 17000 16S rRNA gene sequences per sample), using 97% similarity as the cut-off for a species level OTU (operational taxonomic unit), activated sludge may contain more than 3000 OTUs in one WWTP (19), comparable to the bacterial diversity in a soil sample while the human gut only has a much less diverse microbial community. With the sub-inhibitory concentration of antibiotics in the flocs of activated sludge, plus the high microbial biodiversity, high biomass density (2-50 g/L), the short distance between cells in activated sludge, the extracellular polymeric substance (EPS) matrix, well-mixing conditions in the aeration tank, etc., activated sludge has been proposed as important hotspot for the dissemination of ARGs into environment and consequent exposure to human beings and livestock. In the activated sludge process, the average generation time of bacteria is about six to nine days. That means that there could be more than 600 generations within 10 years of operation in which to develop resistance. Some resistant bacteria in activated sludge may go into the bulk water of the aeration tank and the sedimentation tank, and be discharged together with the effluent.
The resistant bacteria in the sludge could contaminate the soil when activated sludge is applied as a soil conditioner/fertilizer as is being practiced in many counties now. The effluent of WWTPs usually carries bacteria of 105 ~106 cells per litre, continually contributing to the resistance pool in the natural water body and sediment, and even soil if effluent is reused for irrigation. Considering the thousands WWTPs applying activated sludge worldwide, the significance of WWTPs in antibiotics resistance spread cannot be ignored (20), although it may not be as serious as those resistances developed in clinical environments and livestock farms (21). However, compared with studies on resistance genes in other environments, like soil (22), faecal samples (23), studies on resistance genes in activated sludge and wastewater effluent are very limited (24) although it has not been totally overlooked.
Various ARGs, for example, resistance genes to β-lactam (25), tetracyclines (26), have been detected in WWTPs, suggesting that WWTPs have an important role in the dissemination of ARGs in water environments. The ARGs remaining in the treated water are discharged through the effluent into the receiving water bodies and the ARGs in sludge would spread into soil through sludge disposal or land application. Therefore, a precise and comprehensive of knowledge of ARGs profiles in WWTPs is critical for understanding the spread of ARGs in the natural environment.
Metagenomic approach to explore ARGs in WWTPs
In the previous studies, many researchers applied qPCR as the main tool to investigate the profile and quantity of ARGs in different environments. However, qPCR results depend on many factors, including reaction chemistry, PCR machines, the matrix effect of the DNA extract, the skill of the operator, etc. It is not a tool of high comparability which could be used for fair comparison among different laboratories. Since ARGs are a global issue now, especially after the 2013 G8 Summit statement on the joint action to control antibiotic resistance, the science and management societies wish to have more comparable data sets to identify the hotspots of the pollution sources, hot areas/countries having high resistance levels, an approach of high comparability is most welcome at this time as they are suited for such a purpose. With the fast development of next generation sequencing, the metagenomics approach is becoming a widely applied method which is affordable for some routine analyses of ARGs in various environments. Additionally, similarity searching-based annotation may give a more comprehensive profile of ARGs than the qPCR method which is limited by the availability of the primers designed so far. In the past few years, several groups have been working on ARGs surveys by using metagenomics approach (27, 28, 29, 30). Their results revealed the major types of ARGs in WWTPs (27) and the relative resistance levels in activated sludge compared to samples from other environments (31).
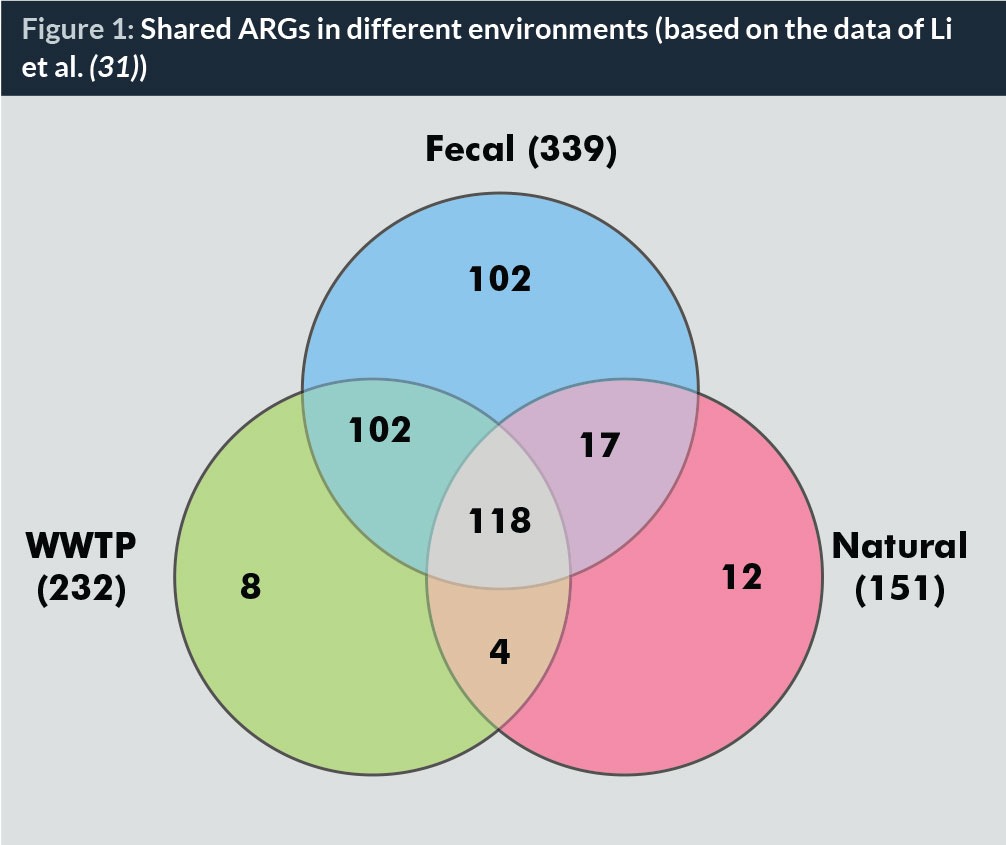
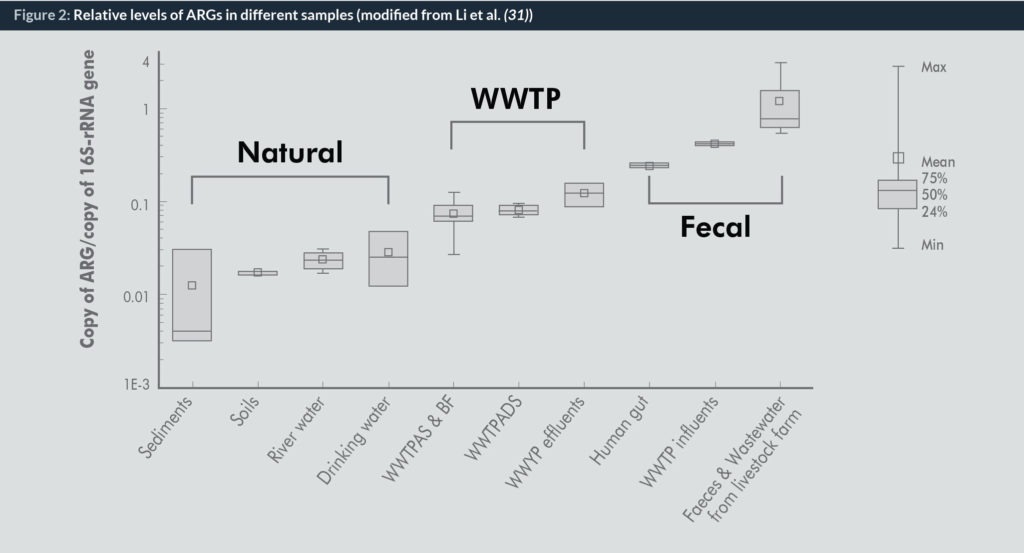
Figures 1 and 2 show the numbers of ARGs types and levels of total ARGs detected in different samples, including faecal samples (animal and human waste), WWTPs and natural environments (sediment, river water, etc.), based on our previous studies (31). A big portion of ARGs in natural environments and faecal samples could be detected in WWTPs (Fig. 1) and the levels of ARGs are comparable to the faecal sources (Fig. 2). Thus, the roles of WWTPs as hotspot for ARGs spread desire further comprehensive study to develop the integrated strategy to fight the resistance.
Using the metagenomic method, Yang investigated the profiles of ARGs in eight activated sludges from a WWTP collected biannually at two seasons (winter and summer) over a four-year period from July 2007 to January 2011, by searching against a structured database of ARGs. The results showed the existence of a broad-spectrum of different ARGs (over 200 subtypes), some of which have never been reported in activated sludge before. The most abundant ARGs were aminoglycoside and tetracycline resistance genes, followed by resistance genes of sulfonamide, multidrug and chloramphenicol. The abundances of some ARG sub-types were inconsistent with that reported in previous studies of activated sludge using the PCR approach. The results of this study demonstrate that a high-throughput-sequencing-based metagenomic approach combined with a structured database of ARGs provides a powerful tool for a comprehensive survey of the various ARGs not only in the activated sludge of a WWTP, but other environmental samples as well. Thus the profiling of ARGs in other ecologically important environmental matrixes may help elucidate those environmental factors contributing to the spread of ARGs. Based on the results about ARGs in a full scale WWTP (32), wastewater influent had the highest ARGs abundance, followed by effluent, anaerobic digestion sludge and activated sludge. Seventy-eight ARGs subtypes persisted through the biological wastewater and sludge treatment process. Anaerobic digestion of sludge may eliminate some ARGs but cannot eliminate all (33, 34).
Metagenomic approaches have also been applied to the investigation of ARGs in other environmental samples. Nesme et al. (29) studied the occurrence and abundance of ARGs in 71 environmental samples. Their results revealed the diverse and abundant ARGs in different environments and suggested these genes were not randomly distributed. Li et al. (31) investigated ARGs and their co-occurrence patterns in 50 samples from 10 typical environments, including WWTPs. In total, 260 ARG subtypes belonging to 18 ARG types were detected with an abundance range of 5.4×10-6 ~ 2.2×10-1 copy of ARG/copy of 16S-rRNA gene for an individual subtype. The trend of the total ARG abundances in different environments matched well with the levels of anthropogenic impacts on these environments. These results demonstrated that WWTPs are important hotspots of ARG dissemination. From the less impacted environments to the seriously impacted environments, the total ARG abundances increased up to three orders of magnitude, i.e., from 3.2×10-3 to 3.1×100 copy of ARG/copy of 16S-rRNA gene. The abundant ARGs were associated with aminoglycoside, bacitracin, β-lactam, chloramphenicol, macrolide-lincosamide-streptogramin, quinolone, sulfonamide and tetracycline, in agreement with the antibiotics extensively used in human medicine or veterinary medicine/promoters.
Perspectives
Now, there is little argument on whether antibiotics and resistance genes should be considered as pollutants in wastewater and treatment plants. However, it is not possible to largely modify the current municipal wastewater treatment infrastructure to achieve high removal efficiency for these pollutants. Then, from engineering and management point of views, what could be done first cost-effectively? A good starting point could be the significant hotspots, like hospital and pharmaceutical wastewater. For hospital wastewater, membrane technology should be used as a pre-treatment before wastewater is discharged into the public sewers connected to municipal wastewater treatment plants, at least an ultrafiltration membrane which could remove viruses and bacteria, or even a nanofiltration membrane which could remove the antibiotics, drugs and free DNA fragments in wastewater effluent. For pharmaceutical wastewater, the biological wastewater treatment process should be discouraged since it is an incubation of superbugs which are discharged into public sewer or natural water body directly. If biological treatment cannot be avoided, a membrane separation unit should be installed to keep the bacteria within the boundary of the pharmaceutical wastewater facilities boundary. The sludge from these hotspots should be incinerated instead of being disposal in other ways, like landfill or land application (no matter for agricultural land or non-agricultural land).
Although there is no doubt now that WWTPs are significant hotspots in spread of ARGs and resistant bacteria, systematic studies are still needed about the fate of antibiotics, resistance genes and resistant bacteria, especially on a global scale with joint efforts. To achieve that target, the scientific and management society require a standardized methodology, like metagenomic approach and high-throughput qPCR. For metagenomic analysis, a more comprehensive and updated database and an open online pipeline are very helpful. With the fast development in metagenomic sequencing, tools (database and annotation pipeline) for the efficient analysis of the huge amount of data are extremely important for timely data processing (35).
Additionally, most of the work on ARGs are based on qPCR assay which is developed based on known ARGs sequences (36) and metagenomics sequencing by referring to the known ARGs in the database (31). However, there could be some unknown ARGs in the environment, especially in the activated sludge system which is poorly understood regarding to ARGs in the microbial cells. To further explore the novel ARGs in wastewater systems, functional screening based on the phenotype should be conducted as demonstrated previously (28).
Furthermore, to fully understand the risk of ARGs from WWTPs, it is important to put the ARGs back to their genetic context, i.e., its host and associated mobile genetic elements. There are some efforts to explore the host of ARGs (37, 38) and mobile genetic elements carrying ARGs (30, 39). However, our current knowledge is still very limited.
Lastly, in recent years, other factors, such as heavy metals (40) and xenobiotic compounds (like triclosan) (41) adsorbed by activated sludge from wastewater bulk water have also been reported as the selective pressures for ARGs via a co-selection mechanism. There is growing concern that contamination of metals may act as selective pressures in the proliferation of antibiotic resistance in wastewater treatment facilities. However, the knowledge on these co-selection factors are very limited and requires further studies in the near future.
As the hotspots, WWTPs have drawn more and more attention regarding its role in resistance dissemination. In addition to the removal of antibiotics in different wastewater treatment process (6, 16, 17, 42, 43), the study on profiles and fate of antibiotic resistance genes in WWTPs could help development of the integrated strategy to fight the resistance and stop ARGs dissemination in the environment (44).
Biography
To download a PDF of this article click here
References
1.Laxminarayan R, Duse A, Wattal C, Zaidi AKM, Wertheim HFL, Sumpradit N, Vlieghe E, Hara GL, Gould IM, Goossens H. 2013. Antibiotic resistance-the need for global solutions. Lancet Infectious Diseases, 13, 1057-1098.
2. WHO. 2014. Antimicrobial Resistance Global Report on Surveillance.
3. Cytryn E. 2013. The soil resistome: The anthropogenic, the native, and the unknown. Soil Biology and Biochemistry. 63, 18-23.
4. Pruden A, Arabi M, Storteboom HN. 2012. Correlation between upstream human activities and riverine antibiotic resistance genes. Environmental Science and Technology. 46, 11541–11549.
5. Ma LP, Li B, Zhang T. 2014. Abundant rifampin resistance genes and significant correlations of antibiotic resistance genes and plasmids in various environments revealed by metagenomic analysis. Applied Microbiology and Biotechnology. 98, 5195-204.
6. Zhang T, Li B. 2011. Occurrence, transformation and fate of antibiotics in municipal wastewater treatment plants. Critical Review on Environmental Science and Technology. 41, 951-998.
7. Kümmerer K. 2009. Antibiotics in the aquatic environment – A review – Part I. Chemosphere. 75(4), 417-434.
8. Li B, Zhang T. 2011. Mass flows and removal of antibiotics in two municipal wastewater treatment plants. Chemosphere. 83(9), 1284-1289.
9. Watkinson AJ, Murby EJ, Kolpin DW, Costanzo SD. 2009. The occurrence of antibiotics in an urban watershed: From wastewater to drinking water. Science of Total Environment. 407(8), 2711-2723.
10. Li B, Zhang T, Xu ZY, Fang HHP. 2009. Rapid analysis of 21 antibiotics of multiple classes in municipal wastewater using ultra performance liquid chromatography-tandem mass spectrometry. Analytica Chimica Acta. 645, 64-72.
11. Batt AL, Bruce IB, Aga DS. 2006. Evaluating the vulnerability of surface waters to antibiotic contamination from varying wastewater treatment plant discharges. Environmental Pollution. 142(2), 295-302.
12. Watkinson AJ, Murby EJ, Costanzo SD. 2007. Removal of antibiotics in conventional and advanced wastewater treatment: Implications for environmental discharge and wastewater recycling. Water Research. 41(18), 4164-4176.
13. Minh TB, Leung HW, Loi IH, Chan WH, So MK, Mao JQ. 2009. Antibiotics in the Hong Kong metropolitan area: Ubiquitous distribution and fate in Victoria Harbour. Marine Pollution Bulletin. 58(7), 1052-62
14. Kasprzyk-Hordern B, Dinsdale RM, Guwy AJ. 2009. The removal of pharmaceuticals, personal care products, endocrine disruptors and illicit drugs during wastewater treatment and its impact on the quality of receiving waters. Water Research. 43(2), 363-380.
15. Lindberg RH, Wennberg P, Johansson MI, Tysklind M, Andersson BAV. 2005. Screening of human antibiotic substances and determination of weekly mass flows in five sewage treatment plants in Sweden. Environment Science & Technology. 39(10), 3421-3429.
16. Li B, Zhang T. 2010. Biodegradation and adsorption of antibiotics in the activated sludge process. Environmental Science and Technology. 44 (9), 3468-3473.
17. Li B, Zhang T. 2013. Removal mechanisms and kinetics of trace tetracycline by two types of activated sludge treating freshwater sewage and saline sewage. Environmental Science and Pollution Research. 20(5), 3024-33.
18. Berendonk TU, Manaia CM, Merlin C, Fatta Kassinos D, Cytryn E, Walsh F, Bürgmann H, Sørum H, Norström M, Pons MN, Kreuzinger N, Huovinen P, Stefani S, Schwartz T, Kisand V, Baquero F, Martinez JL. 2015. Tackling antibiotic resistance: the environmental framework. Nature Reviews Microbiology. 13(5), 310-7.
19. Zhang T, Shao MF, Ye L. 2012. 454 Pyrosequencing reveals bacterial diversity of activated sludge from 14 sewage treatment plants. ISME Journal, 6, 1137-1147.
20. Czekalski N, Díez EG, Bürgmann H. 2014. Wastewater as a point source of antibiotic-resistance genes in the sediment of a freshwater lake. ISME Journal. 8, 1381–1390.
21. Munck C, Albertsen M, Telke A, Ellabaan M, Nielsen PH, Sommera MOA. 2015. Limited dissemination of the wastewater treatment plant core resistome. Nature Communication. 6, 8452.
22. Howe AC, Jansson JK, Malfatti SA, Tringe SG, Tiedje JM, Brown CT. 2014. Tackling soil diversity with the assembly of large, complex metagenomes. PNAS. 111, 4904–4909. 10.1073/pnas.1402564111.
23. Looft T, Johnson TA, Allen HK, Bayles DO, Alt DP, Stedtfeld RD. 2012. In-feed antibiotic effects on the swine intestinal microbiome. PNAS. 109(5), 1691-1696.
24. Zhang XX, Zhang T, Fang HHP. 2009. Antibiotic resistance genes in water environment. Applied Microbiology and Biotechnology. 82(3), 397-414.
25. Yang Y, Zhang T, Zhang XX, Liang DW, Zhang M, Gao DW, Zhu HG, Huang QG, Fang HHP. 2012. Quantification and characterization of β-lactam resistance genes in 15 sewage treatment plants from East Asia and North America. Applied Microbiology and Biotechnology. 95(5), 1351-1358.
26. Zhang T, Zhang M, Zhang XX, Fang HHP. 2009. Tetracycline resistance genes and tetracycline resistant lactose-fermenting Enterobacteriaceae in activated sludge of sewage treatment plants. Environmental Science and Technology. 43, 3455–3460.
27. Yang Y, Li B, Ju F, Zhang T. 2013. Exploring variation of antibiotic resistance genes in activated sludge over a four-year period through a metagenomic approach. Environmental Science and Technology. 47(18), 10197-10205.
28. Forsberg K, Reyes A, Wang B, Selleck EM, Sommer MOA, Dantas G. 2012. The shared antibiotic resistome of soil bacteria and human pathogens. Science. 337, 1107-1111.
29.] Nesme, J, Cecillon, S, Delmont, TO, Monier, JM, Vogel, TM, Simonet, P. 2014. Large-scale metagenomic-based study of antibiotic resistance in the environment. Current biology: CB. 24, 1096-1100.
30. Li AD, Li LG, Zhang T. 2015. Exploring antibiotic resistance genes and metal resistance genes in plasmid metagenomes from wastewater treatment plants. Frontiers in Microbiology. 6, doi: 10.3389/fmicb.2015.01025.
31. Li B, Yang Y, Ma LP, Ju F, Guo F, Tiedje JM, Zhang T. 2015. Metagenomic and network analysis reveal wide distribution and co-occurrence of environmental antibiotic resistance genes. ISME Journal. 9(11), 2490-502.
32. Yang Y, Li B, Zou SC, Fang HHP, Zhang T. 2014. Fate of antibiotic resistance genes in sewage treatment plant revealed by metagenomic approach. Water Research. 62, 97-106.
33. Ju F, Li B, Ma LP, Wang YB, Huang DP, Zhang T. Antibiotic resistance genes and human bacterial pathogens: co-occurrence, removal, and enrichment in municipal sewage sludge digesters. Water Research. 91, 1-10.
34. Zhang T, Yang Y, Pruden A. 2015. Effect of Temperature on removal of antibiotic resistance genes by anaerobic digestion of activated sludge revealed by metagenomic approach. Applied Microbiology and Biotechnology. 99(18), 7771-9.
35. Yang Y, Jiang XT, Zhang T. 2014. Evaluation of a hybrid approach using UBLAST and BLASTX for metagenomic sequences annotation of specific functional genes. PLoS ONE. DOI: 10.1371/journal.pone.0110947.
36. Zhang T, Zhang XX, Ye L. 2011. Plasmid metagenome reveals high levels of antibiotic resistance genes and mobile genetic elements in activated sludge. PLoS ONE. 6(10), e26041.
37. Ma LP, Xia Y, LI B, Yang Y, Li LG, Tiedje JM, Zhang T. 2016. Metagenomic assembly reveals hosts of antibiotic resistance genes and the shared resistome in pig, chicken and human feces. Environmental Science and Technology. 50, 420-427.
38. Li B, Zhang XX, Guo F, Wu WM, Zhang T. 2013. Characterization of tetracycline resistant bacterial community in saline activated sludge using batch stress incubation with high-throughput sequencing analysis. Water Research. 47(13):4207-16.
39. Zhang XX, Zhang T, Zhang M, Fang HHP, Cheng SP. 2009. Characterization and quantification of class 1 integrons and associated gene cassettes in sewage treatment plants. Applied Microbiology and Biotechnology. 82(6), 1169-77.
40. Seiler C, Berendonk TU. 2012. Heavy metal driven co-selection of antibiotic resistance in soil and water bodies impacted by agriculture and aquaculture. Frontiers in Microbiology. 3:399. doi: 10.3389/fmicb.2012.00399.
41. Pal C, Bengtsson-Palme J, Kristiansson E, Larsson DGJ. 2015. Co-occurrence of resistance genes to antibiotics, biocides and metals reveals novel insights into their co-selection potential. BMC Genomics. 16, 964.
42. Li B, Zhang T. 2012. pH significantly affects removal of trace antibiotics in chlorination of municipal wastewater. Water Research. 46, 3703-3713.
43. Li B, Zhang T. 2013. Different removal behaviours of multiple trace antibiotics in municipal wastewater chlorination. Water Research. 47(9), 2970-82.
44. Davies J. 1994. Inactivation of antibiotics and the dissemination of resistance genes. Science, 264, 375-382.