Professor Ellen Silbergeld, Johns Hopkins University, Bloomberg School of Public Health, USA; Dr Awa Aidara-Kane, Coordinator WHO–Agisar and Jennifer Dailey, Phd Student in Materials Science and Engineering, Johns Hopkins University, USA
All uses of antimicrobials contribute to the emergence and dissemination of resistance, and therefore effective interventions to control antimicrobial resistance (AMR) must address all of the major uses of antimicrobials in both healthcare and agriculture. There is ample evidence that AMR can emerge and be disseminated from agriculture to human populations. This evidence includes first reports of resistance, increases and decreases in AMR in human isolates following additions and bans on antimicrobial use in animal feeds, and resistance associated with first use of novel molecules in agriculture prior to clinical use. In terms of amplification, agricultural practices are more likely to support global spread and persistence of AMR in the microbiome. All uses of antimicrobials should be evaluated in an integrated manner because of the interconnectedness of agriculture and human health, particularly through the food supply. This interconnectedness makes it impossible and potentially misleading to attempt to control and separately estimate the attributable risk of each source.
The debate over the importance of antibiotic use in agriculture as a driver of antimicrobial resistance (AMR) has been active since the first reports of drug resistance in agricultural settings, which coincided with the first reports of drug resistance in clinical settings. In order to move forward in developing concerted programmes at local, national and global levels for responding to the crisis in AMR, we need to incorporate state-of-the-art scientific understanding of molecular microbiology as well as the state-of-the-science on AMR selection and dissemination.
There are very few differences between the conditions of antimicrobial use in agriculture and in healthcare in terms of drivers for the emergence of resistance. This is not surprising: many of the conditions under which antimicrobials are used in food animal production – overcrowding, lack of sanitation, administration of drugs without information on target pathogens or antimicrobial susceptibility, failure to treat wastes prior to discharge – are the same as the risk factors that we have long recognized as drivers of AMR in human healthcare settings. However, unlike healthcare, there are few programmes or regulations to ameliorate these conditions. Other conditions of antimicrobial use are specific or more common to agriculture – subtherapeutic exposures (below the MIC); extended duration of dosing; co-exposures to multiple antimicrobials administered in feeds, lack of infection control and incomplete biosecurity and biocontainment throughout the food production chain (1). These conditions amplify the dissemination and persistence AMR, including multidrug-resistant strains of zoonotic pathogens and they ultimately contribute to the major reservoirs for AMR genes in animal guts and in the environment. On a mass basis, the major route of release from food animal production to the environment is through disposal of untreated animal wastes from farms and processing plants. In addition to AMR-resistant pathogens and resistance genes, these wastes contain unmetabolized antimicrobials and spilled feeds containing antimicrobials. As a result, soils, sediments, and surface waters in agricultural areas are contaminated by AMR genes, antimicrobials and AMR bacteria. These environmental microbiomes are the ultimate reservoirs of AMR (including resistomes or the microbial pangenomic resources for resistance and mobilomes or the resources for gene transfers among microbes) (2).
Mechanisms of AMR
Antimicrobials are natural products and AMR has existed for billions of years within and among microbial communities as “bioweapons” used by microbes in competition for nutrients and space. This warfare has resulted in pressure for survival of these attacks through evolutionary processes that select for traits and genes that confer resistance to these natural toxins. Pressures from human uses of antimicrobials are very recent, following on not only their discovery and isolation but also the development of mass production in the 1940s and marketing for both clinical medicine and agriculture.
During the first years following discovery and production of antimicrobials, this prehistory of AMR was largely unknown. As a result, it was generally thought that AMR depended solely on evolutionary mechanisms for the emergence of antibiotic resistance. This is a random process, highly dependent upon growth rates of bacterial populations. Studies of bacterial response to antimicrobial pressure identified an additional mechanism independent of growth rate and genetic mutation that involved horizontal transfers of resistance genes among bacteria (3). Later research demonstrated that these horizontally transferred genes could be incorporated into the genome of the recipient cell, and the cell was able to synthesize protein targets of drugs that were altered such that these targets were resistant to the same drugs. We now recognize that horizontal gene transfer (HGT), rather than mutation, is the major mode by which bacteria respond to antimicrobial pressure through multiple mechanisms including conjugation or exchange through direct cell:cell contact, transformation or incorporation of naked DNA from the extracellular environment, and transduction or horizontal gene transfer (4). Because HGT does not depend upon the random events of evolutionary adaptation, the “cost of resistance” theory does not apply. This theory holds that in the absence of a stressor such as antimicrobials, the microbial population reverts to the “wild” type since the presence of the gene mutation must exert some cost in terms of energetics or reproductive rates (5). The dominant role of HGT explains why AMR may persist after removal of drug pressures. HGT also transfers individual resistance genes as well as cassettes of multiple genes that encode for co-resistance and co-selection of resistance within and among microbial communities.
We still have important gaps in our knowledge of the mechanisms of AMR. Many of these introduce bias towards underestimating the problem. For example, we continue to focus on examining AMR in pathogenic bacteria, a definition that is relevant to human health but not necessarily to the networks of gene exchange within the microbial world or microbiome including commensal strains. Within the environment and animal guts, most of the resistome is present in commensal strains (6, 7). Also in general we define resistance phenotypically, that is, by the response of bacterial cultures to in vitro antimicrobial challenge. We do not assess the presence of resistance genes within specific bacterial strains or the microbial community (or microbiome). In terms of understanding and controlling AMR, we need to assess the entirety of the resistome and the mobilome (2).
Conditions of AM use in agriculture as risk factors for emergence and dissemination
Food animal production has been identified as a source of human infections by virulent and drug-resistant infections as early as studies of farmers by Williams Smith (8). We need to understand and recognize the similarities and differences in antimicrobial use in agriculture and healthcare. Overall, the most significant risk factor driving AMR emergence in any setting is the volume of drug use, as recognized by Fleming in his Nobel Prize speech (9). Empirical data supports this for both clinical and agricultural uses in studies of both uses and bans (10). In many countries, the largest use of many antimicrobials is in agriculture, not medicine (11). The response of the microbiome to antimicrobial pressure is shaped by the characteristics of antimicrobials exposure, which often differ between agriculture and healthcare. Subtherapeutic exposures, which are common in food animal production, are particularly effective as drivers of selection for AMR by driving cell division and mutation as well as horizontal transfer of resistance genes. Continuous or prolonged low- level antimicrobial use expands resistomes in hosts and the environment. Multidrug pressure, also common in food animal production, enhances the emergence and dissemination of multidrug resistance though horizontal transfer of multiple resistance genes (12).
Other practices and policies in agriculture have also contributed to the erosion of antimicrobial effectiveness in medicine, including the approval of new antimicrobials for agricultural use prior to registration for clinical medicine (13-15) as well as approval of a clinical drug for agricultural use, which greatly increases the overall volume of antimicrobial use. Studies from many countries have demonstrated large increases in the prevalence of quinolone resistance in human isolates of Campylobacter after approval of enrofloxacin, a quinolone drug, for use in poultry feeds (16).
Dissemination

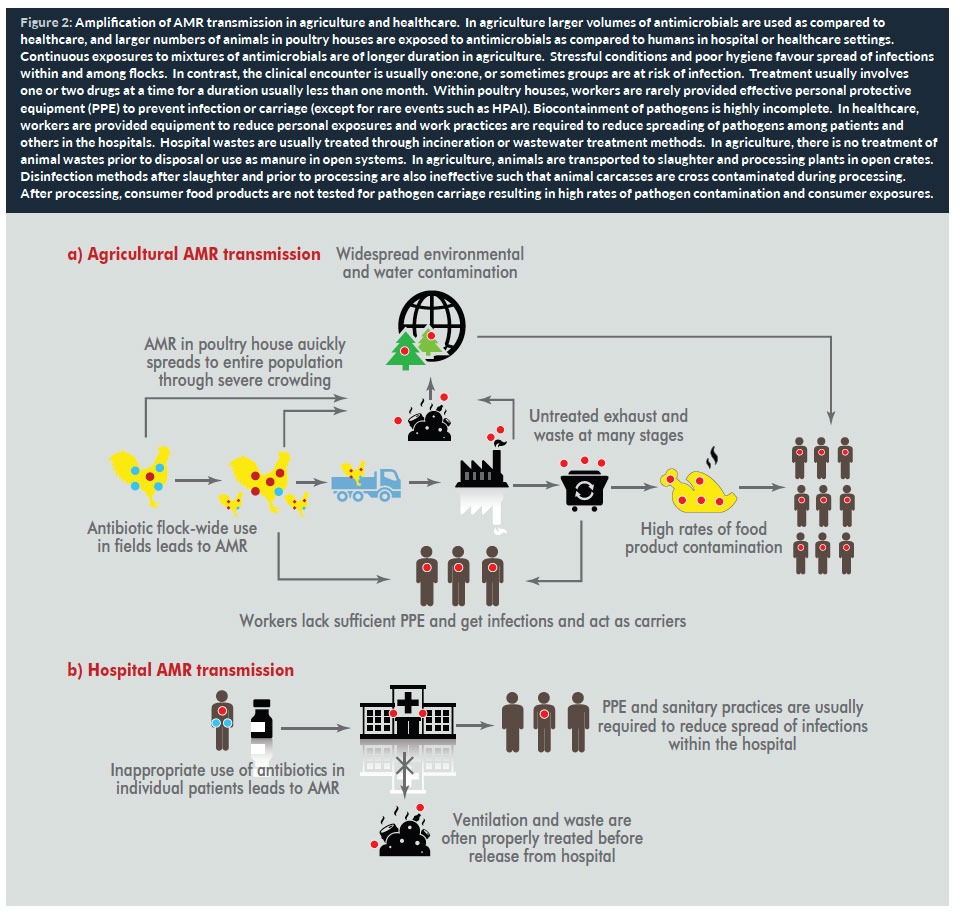
The most significant risk factor for AMR dissemination from animals to humans is through the food supply. We recognize the importance of food products from animals, but another important but often overlooked pathway of dissemination is through the environment to food crops consumed by humans. Examples have involved crops grown with animal wastes (as in organic production) or grown with irrigation by surface water sources contaminated by run off from land disposal of animal wastes. Food animal production is a multi-step process that amplifies the prevalence of resistance and opportunities for dissemination among animals and human populations (17). This is shown in Figure 2 for agriculture (a) and hospitals (b).
Conditions of antimicrobials use in agriculture
The conditions of antimicrobials use in agriculture include the major drivers of resistance and dissemination: large volumes of antimicrobial use, sub-therapeutic and extended exposures of animal gut microbiomes, co-exposures to drug mixtures and other agents in feeds (including metals), regional and international movement of animals and food products, production methods in agriculture, and exposures of workers and other biota. AMR zoonotic pathogens and AMR genes have been detected in food animals and their wastes, in carcasses and consumer food products, and in air and wastewater at retail outlets such as wet markets. Among these pathogens, there is an extensive literature on the presence of AMR strains of Staphylococcus aureus, Escherichia coli, Enterococcus faecalis, and a range of Gram-negative pathogens in all these settings (18, 19).
Extended spectrum β lactamase (ESBL)+ in Enterobacteriaceae is an exemplary case of the importance of an integrated perspective uniting agriculture, food, and the environment. ESBL+-strains of many of highly pathogenic bacteria have now been reported worldwide from the Americas, Europe, Asia-Pacific, and Africa (20). Carbapenemase-producing strains of Klebsiella pneumoniae, Escherichia coli, Pseudomonas aeruginosa, Acinetobacter baumannii and other pathogens are also increasingly reported in isolates from human populations. Almost as soon as new extended spectrum β lactam antimicrobials have been developed and approved for use in healthcare, they have also been permitted for use in food animal production in many countries (21). Not surprisingly, reports of new ESBL+ strains are also quickly reported from both healthcare and agriculture. Although carbapenems are not generally registered for use in agriculture, carbapenemase genes have in bacteria carried by food animals and other biota, possibly driven by over use of extended spectrum cephalosporins (22). These drugs are used without formal approval in other regions. A recent study of smallholders in Peru reported on resistance to cephalosporins and many other antimicrobials in E. coli isolates from poultry and farmers (23).
We recognize that contamination of the food supply is a critically important event for the generalized dissemination of resistant pathogens from agriculture to human populations. For that reason, there are national programmes for monitoring the presence of zoonotic pathogens and AMR strains in food products. There are fewer programmes for reporting on antimicrobial use in agriculture. Very few of these are systematic and most of these do not include cephalosporins. However, the environment is equally important as a reservoir for resistance genes primarily through releases throughout the stages of food production and the disposal of untreated animal wastes on land. Despite the importance of the environment in containing AMR, there are no systems of environmental monitoring of AMR so that we cannot fully estimate the current magnitude of these routes of exposure. There are studies from several regions that have reported recent increases in the presence of ESBL genes and resistance in pathogens isolated from poultry and other food animals, as well as from food products and within the environment of animal slaughter and processing plants (24). Moreover, to an increasing extent, ESBL resistance is coupled with other resistance determinants, including cystins, packaged in transferrable gene cassettes.
Cross movement between agriculture and healthcare
The most important issue in resolving the continuing debate over the relative importance of agriculture and healthcare as sources of AMR is to understand that these two sectors are not independent. The same pathways – food and environment — are common to both healthcare and agriculture, since both food and people move between these settings. There are no real barriers between the movement of the same strains of AMR pathogens between agriculture and healthcare settings. This movement goes both ways as shown in a well annotated genomic history of the cross transmission of so-called “livestock” strains of MRSA (ST398) from humans to animals and from animals to humans (25). These interactions make it difficult, if not impossible, to define and quantify agricultural- and healthcare-related attributable risks of human exposure to AMR pathogens.
Conclusions
There is adequate evidence to prioritize agricultural use of antimicrobials as sources of exposure of human populations to ESBL+ Enterobacteriaceae and other AMR zoonotic pathogens. The reservoirs for AMR genes include animal and human host guts, animal wastes, and the environments into which releases from animal confinement houses and wastes are discharged without pretreatment or monitoring. The molecular mechanisms driving AMR emergence and dissemination in agriculture involve conditions known to drive selection for resistance through stimulation of both evolutionary and non-evolutionary mechanisms. These are in large part the same conditions that have long been recognized as major risk factors for AMR in human healthcare settings. Unlike healthcare, there are no programmes to ameliorate these conditions. Other conditions of antimicrobial use in agriculture – sub-MIC exposures; long duration dosing; multiple agents administered in feeds, lack of waste management and lack of biosecurity and biocontainment – favour emergence, persistence and dissemination of AMR, including MDR phenotypes.
Much of the debate on this topic is misplaced. Because of the interconnectedness of agriculture and human populations, it is not likely to be possible to estimate the attributable risk of agricultural and non-agricultural sources of AMR. We should be more concerned that our information on sources and prevalence of AMR is skewed since our knowledge of settings other than healthcare and nonpathogenic organisms is almost non-existent.
Improvements in both surveillance and monitoring of agriculture (including food, workers on farms and in slaughter and processing) as well as the environment are essential for identifying critical opportunities for early detection and the development of interventions specific to agricultural antimicrobial use as part of the overall WHO programme on responding the threat of AMR. This is not to disregard the importance of ensuring prudent use of antimicrobials in healthcare; this article demonstrates the importance of developing guidance and programmes that extend to and integrate findings on all uses and abuses of AMs.
Support: Research in Dr Silbergeld’s laboratory on this topic has been supported by US National Institutes of Health and Centers for Disease Control as well as the Pew Charitable Trust. The authors thank many students and colleagues at Johns Hopkins and globally for their contributions to this research, as well as members of the Advisory Group on International Studies of Antimicrobial Resistance (WHO) for their insights.
Biographies
Dr Awa Aidara-Kane, MSc, PhD is involved in laboratory capacity-strengthening activities for the Global Foodborne Infections Network (GFN) and leads World Health Organization (WHO) activities on the containment of antimicrobial resistance (AMR) from a food safety perspective. She is Coordinator of the WHO Advisory Group on the Integrated Surveillance of Antimicrobial Resistance (WHO-AGISAR) and the WHO focal point for FAO/OIE/WHO Tripartite activities on AMR. She also leads the WHO One Health workstream for the implementation of the Global Action Plan on AMR. Before joining WHO in 2002, she was Associate Professor of Bacteriology and Virology in Senegal from 1984 to 1990, and Researcher at the Pasteur Institute in Dakar from 1985 to 2002. At Institute Pasteur, she was Head of the Laboratory of Research in Bacteriology and Scientific Director of the Laboratory of Food Hygiene and Environment. Her area of research was the characterization of virulence factors and AMR determinants in enteric and foodborne pathogens. She holds a MSc in Food Technology and a PhD in Microbiology from the University of Technology of Compiegne, France.
Jennifer Dailey, BA, MSE, is a PhD student in Materials Science and Engineering at the Whiting School of Engineering at Johns Hopkins University. Her research interests involve increasing food security through the creation of enhanced biosensors based on organic electronic devices and pH-reactive polymers. She is also interested in point-of-care vapour sensors to help track environmental pollutants. She received her BA in Physics from the University of Pennsylvania in 2012 and her MSE in Materials Science and Engineering from Johns Hopkins in 2015.
References
1. Leibler JH, Dalton K, Pekosz A, Gray GC, Silbergeld EK. Epizootics in Industrial Livestock Production: Preventable Gaps in Biosecurity and Biocontainment. Zoonoses Public Health. 2016 64: 137-145
2. Gillings MR. Evolutionary consequences of antibiotic use for resistome, mobilome, and microbial pathogens. 2013 Front Microbiol 4:4
3. Tatum EL and Lederberg J. Gene recombination in the bacterium Escherichia coli 1947 J Bacteriol. 1947 53(6):673-84.
4. Von Wintersdorff CJH, Penders J, van Niekerk JM et al. Dissemination of antimicrobial resistance in microbial ecosystems through horizontal gene transfer. 2016 Front Microbiol 7:173
5. Levin BR, Concepcion-Acevedo, and Udekwu KI. Persistence: a copacetic and parsimonious hypothesis for the existence of on-inherited resistance to antibiotics. 2014 Curr Opin Microbiol 18-21.
6. Shtertzer N and Mizrahi I. The animal gut as a melting pot for horizontal gene transfer. 2015 Can. J. Microbiol. 63: 603-605
7. Ravi A, Avershina E, Foley SL et al. The commensal infant gut meta-mobilome as a potential reservoir for persistant multidrug resistance integrons. 2015 Nature Scientific Rep 5:15317
8. Williams Smith H and Crabb W E. The effect of diets containing tetracyclines and penicillin on th Staphylococcus aereus flora of the nose and skin of pigs and chickens and their human attendants. 1960 J Pathol Bacteriol 79: 343
9. Fleming, Alexander- Banquet Speech. Nobelprize.org. Nobel Media AB 2014. Web. 2 Apr 2017. http://www.nobelprize.org/nobel_prizes/medicine/laureates/1945/fleming-speech.html
10. Hammerum AW, Heuer OE, Emborg H D et al. Danish integrated antimicrobial resistance monitoring and research program. 2007. Emerg Infect Dis 13: 1632-9.
11. Van Boeckel TP, Brown C, Gilbert M, et al. Global trends in antimicrobial use in food animals. 2015 PNAS 112: 5649-4654.
12. ter Kuile BH, Kraupner N, and Brul S. The risk of low concentrations of antibiotics in agriculture for resistace in human health care. 2016 FEMS Microbiol Let 363: fnw210
13. Smith DL, Harris AD, Johnson JA, Silbergeld EK, Morris JG Jr. Animal antibiotic use has an early but important impact on the emergence of antibiotic resistance in human commensal bacteria. Proc Natl Acad Sci USA. 2002 Apr 30;99(9):6434-9
15. Kieke AL, Borchardt MA, Kieke BA, et al. Use of streptogramin growth promoters in poultry and isolation of streptogramin-resistant Enterococcus faecium from humans. 2006 J Infect Dis 194: 1200-1208.
15. Wittum TE. The challenge of regulating agricultural ceftiofur use to slow the emergence of resistance to extended spectrum cephalosporins. 2012 AEM 28: 7819-7821
16. Angulo FJ, Baker NL, Olsen SJ et al. Antimicrobial use in agriculture: controlling the transfer of antimicrobial resistance to humans. 2004 Sem Ped Infect Dis 15:78-85
17. Berendonk TU, Monaia CM, Merlin C, et al. Tackling antibiotic resistance: the environmental framework. 2015 Nature Reviews 13: 310
18. Silbergeld EK, Davis M, Leibler JH, Peterson AE. One reservoir: redefining the community origins of antimicrobial-resistant infections. 2008 Med Clin North Am. 92:1391-407
19. Skurnik D, Clermont O, Guillard T, et al. Emergence of antimicrobial resistant Escherichia coli of animal origin spreading in humans. 2015 Mol Biol Evol 33: 898-914.
20. Molton J, Tambyah PA, Aug BSP et al. The global spread of healthcare-associated multidrug-resistant bacteria: a perspective from Asia. 2013 Clin Infect Dis 56: 1310
21. Greko C [for the Scientific Advisory Group on Antimicrobials for Medicinal Products and Veterinary Use]. Reflection paper on the use third and fourth-generation cephalosporins in food producing animals in the European Union. 2009 J Vet Pharmacol Therap 32:515-533.
22. Mollenkopf DF, Stuff JW, Mathys DA et al. Carbapenemase-producting Enterobacteriaceae recovered from the environment of a swine farrow-to-finish operation in the US. 2016 Antimicrob Agents Chemotherap 61(2). pii: e01298-16
23. Braykov NP, Eisenberg JNS, Grossman M, et al. Antibiotic resistance in animal and environmental samples associated with small-scale poultry farming in northwestern Ecuador. 2016 mSphere1:e00021-15
24. Leverstein-van Hall MA, Dierikx CM, Cohen Stuart J et al. Dutch patients, retail chicken meat and poultry share the same ESBL genes, plasmids, and strains. 2011 Clin Microbiol Infect 17: 873-880
25. Price LB, Stegger M, Hasman H et al. Staphylococcus aureus CC398: host adaptation and emergence of methiciliin resistance in livestock. 2012 mBio 3:e00305-11.