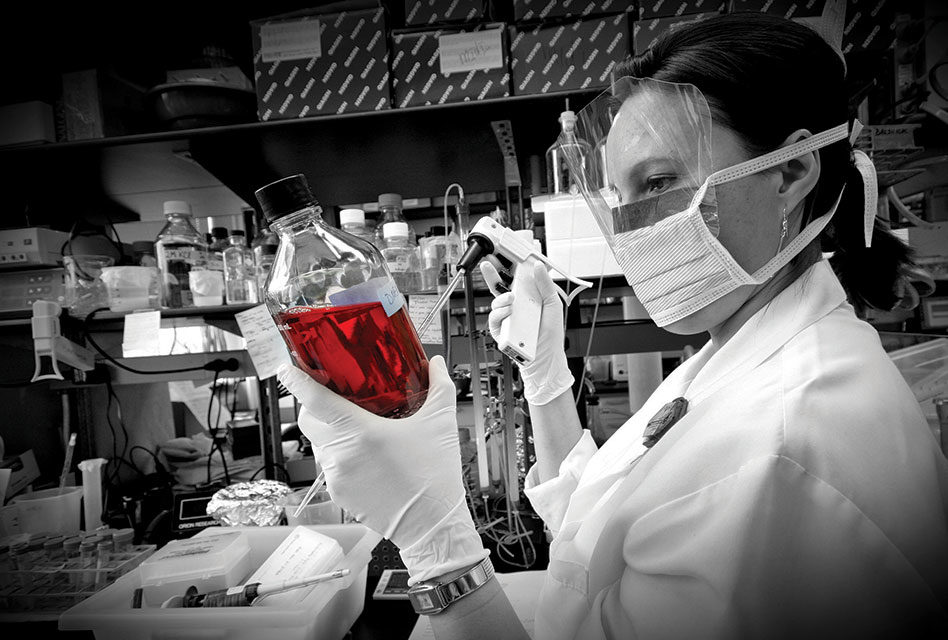
Soumya Swaminathan (left), Secretary, Department of Health Research (Ministry of Health & Family Welfare), Government of India and Director General, Indian Council of Medical Research (ICMR) and Kamini Walia (right), Division of Epidemiology and Communicable Diseases, ICMR, New Delhi, India.
The development of resistance in pathogens of public health importance is getting the attention of all relevant stakeholders. It is clear that the challenge of antimicrobial resistance (AMR) is assuming alarming proportions and will be a biggest killer in years to come if a timely and effective response is not put forward to address this challenge. It is now well understood that antimicrobial resistance is a multifaceted challenge and unless we address all the aspects of the issue the response will not be adequate. Research and development in antimicrobial resistance in the last few decades has only focused on the development of new drugs. Although availability of good quality diagnostics and socioeconomic aspects have been recognized as major contributors to the increasing burden of antimicrobial resistance, very little has been done to develop new diagnostics or understand social, behavioural and economic factors that lead to drug use or misuse. Similarly, understanding of the burden of antimicrobial resistance, the pathogenesis and host genetics was not linked to the development of new drugs and interventions due to lack of interest from the pharmaceutical industry, which has steered clear of antimicrobial research. This article discusses the ongoing research and development initiatives and highlights the need for effective convergence of these efforts, such that the data is interpreted in a more holistic manner leading into the development of preventive and treatment interventions which are relevant, affordable and accessible.
Antibiotics and antibiotic resistance are not modern phenomena. The problem of antibiotic resistance existed subsequent to the discovery of the first antibiotic, Penicillin, in 1928, with resistance to Penicillin reported as early as 1940. Resistance to other antimicrobial agents, such as streptomycin, sulphonamides, beta-lactams, tetracyclins, microlides and quinolones was reported within a decade or less of their discovery (1). It is the combination of various factors like negligent broad-spectrum antibiotic prescription practices among medical practitioners, non-compliance to the recommended treatment duration and non-judicious use of antimicrobial agents by populations which is driving the emergence of resistance in infectious agents of public health relevance. The resistant strains hence generated spread to a larger population either from the community or through the hospital-acquired route (2). Antimicrobial resistance leads to serious escalations in healthcare costs and this burden is still poorly quantified, globally. As per the recent estimates, antimicrobial resistance, if allowed to grow unchecked, will cost the world an additional 10 million lives a year by 2050, with a cumulative cost of US$ 100 trillion, more than one and a half times the annual world GDP today (3).
While antimicrobial resistance is a problem with both developed and developing countries alike, countries like India are considered as one of the epicenters of this problem for multiple reasons. India has a large burden of infectious diseases and the treatment of infections in India is often mishandled owing to many factors, including lack of appropriate diagnostic support in terms of quality and availability, absence of outbreak reports by any national agency for wider dissemination and the tendency of doctors to continue age old prescription practices, as well as various socioeconomic factors (2). The management of many bacterial infections, often presenting as main syndrome fever, remains empirical. In India, and many other low- and middle-income countries (LMICs,) weak antibiotic policies, coupled with poor antibiotic resistance surveillance, leads to a build-up of antibiotic resistance that contributes to the global pool of difficult-to-treat-infections (4).
The traditional approach to combat infectious diseases and antimicrobial resistance has been reliance on new drugs and drug combinations as seen in TB and malaria programmes. The pharmaceutical industry has so far been able to provide us with alternatives which have helped the medical fraternity to treat patients. But the picture ahead is not very promising as the pipeline of antibiotics is fast drying up and no new antibiotic molecule has been reported since 1987 (5). The increase in consumption of antibiotics like colistin and carbapenems reflects the grim picture of running out of treatment options which is forcing us to look for alternatives to salvage the situation as it exists on the ground today. The discussion on the measures to reduce antimicrobial resistance lists restricting antimicrobial use as one of the top priorities. However, restricting use alone may not be the answer in LMICs where still more children die from lack of access to treatment than from drug-resistant bacteria.
In order to counter the threat that antimicrobial resistance poses, we need to strengthen our understanding at multiple levels so that we are able to launch an informed response in each of the gap areas. Firstly, there is a need to have a correct estimate of the drug-resistant infections for which effective and responsive surveillance systems are needed at national and regional levels. Secondly, rapid and specific diagnostics are needed to reduce unnecessary prescription and improve the quality of prescribing in situations where necessary. Thirdly, it is important to create tools which are guided by research on sociobehavioural issues, to create awareness among prescribers and communities. Fourthly, there is a need to develop new treatment options by supporting research on new drug molecules and drug targets and repurposing old antibiotics. Lastly, all this effort will be futile if we do not create systems to enable judicious utilization of what we have and reduce infections in hospitals so that there are fewer situations warranting antibiotic use. This article will look at the progress made in addressing each of these challenges through research and innovation in order to find plausible solutions to address the challenge of antimicrobial resistance in the years to come.
Strengthening surveillance
The generation of reliable, relevant and up-to-date data on antimicrobial resistance is most important to build an effective response. This makes establishing laboratory-based surveillance of antibiotic resistance a fundamental priority for all the countries. In the face of rising resistance, some countries have responded by establishing national surveillance networks, while others have not. The World Health Organization (WHO) recently launched The Global Antimicrobial Resistance Surveillance System (GLASS) to support a standardized approach to the collection, analysis and sharing of data on antimicrobial resistance at a global level, in order to inform decision-making, drive local, national and regional action and provide the evidence base for action and advocacy. GLASS aims to combine clinical, laboratory and epidemiological data on pathogens that pose the greatest threats to health globally. The GLASS manual details the proposed approach for the early implementation of the surveillance system that will focus on antibiotic-resistant bacteria, and outlines the flexible and incremental development of the system over time that will incorporate lessons learned from the early implementation phase (6).
In India, the Indian Council of Medical Research (ICMR) has recently set up an antimicrobial resistance surveillance network, which is currently networking the tertiary care hospitals in the country to collect nationally representative data with an aim to providing evidence which can guide treatment practices in the country. From the data obtained so far, more than 70% of Enterobacteriaceae are resistant to third-generation cephalosporins. Among the Enterbacteriaceae species, Klebsiella and E.coli have been found to be resistant to third-generation cephalosporins (80%). Many hospital-acquired infections are caused by Acinetobacter baumanii and Pseudomonas aeruginosa, showing 50% resistance to carbapenems. The data emanating from this network since the last quarter of 2013 has shown that S.typhi multidrug resistance (MDR) to ampicillin, chloramphenicol and trimethoprim/sulfamethoxazole is showing a downward trend and resistance to fluoroquinolones/cephalosporins in S. typhi is increasingly reported. This finding is very promising as this provides evidence to start using first-generation, simple, cheap drugs. (ICMR data).
There are still many gaps in our understanding of the molecular and cellular biology of bacterial resistance, especially how bacteria evolve, acquire and transmit antibiotic resistance and how they adapt to life in human and animal hosts. In addition to strengthening the surveillance at regional and national level, there is need to understand the mechanisms of drug resistance at the molecular level, which will eventually provide guidance for and also lead to generation of knowledge which can guide development of new diagnostics, potential drug targets and drug molecules. Advances in genomics, systems biology and structural biology have enabled greater understanding of the events and mechanisms leading to resistance. This includes understanding the role of porins, efflux pumps and synergistic mechanisms that lead to drug resistance in manners different from the way it has been conventionally understood. This advancement has led to the discovery and development of new potential drug molecules and approaches that can negotiate existing resistance mechanisms, as well as new targets. Hence, we have the ability to forecast the emergence of resistance to novel drugs, locate exactly where and when this might occur and delineate the mechanisms responsible. The timely understanding of naturally occurring resistance mechanisms and targets that can allow numerous structural alterations should lead to the discontinuation of the development of agents that are likely to fail in the clinic as a result of resistance. An early stage of drug development, if supported by scientific evidence from studies on resistance development and mechanisms of resistance, will also facilitate the development of dosing regimens that can help to minimize the emergence of resistance to current and new antibiotics, enabling these drugs to be used to best effect and focus resources on agents that are less likely to fail (7).
Improving diagnosis
Absence of a reliable, affordable and rapid point of care diagnostic test is central to the problem of antimicrobial resistance, which is being fuelled by unnecessary antibiotic use. Empirical decision-making, in the absence of confirmed diagnosis, results in the patient getting the treatment quickly in LMICs, but is also a major driver of resistant infections and treatment failures. A point of care test to rapidly and accurately differentiate between viral and bacterial infections, like a much improved version of serum procalcitonin or molecular signature patterns, is a priority for LMICs, where the diagnostic capacities are limited and antibiotic prescriptions are routinely given out for viral prescriptions as there is no way to conclusively differentiate bacterial infections from viral. However, there are many issues, such as how many pathogens to be included in the rapid test, whether the test should focus on identification of pathogen or only the drug resistance, and should the test also define molecular load to differentiate between colonization and infection. There is a need to create a consensus on these technical dilemmas to facilitate progress on development of a POC. Since most rapid tests use a microfluidic platform, a microscale assay that provides integrated functionalities in a minimum of unit steps needs to be designed to replace the conventional benchtop sample preparation in situations with no laboratory access. Consensus on the best sampling sites for detection of pathogens in many infectious diseases is also needed (8).
Among LMICs such as India, use of broad spectrum antibiotics has led to development of multidrug resistance (including to beta-lactams and carbapenems) among the gram negatives and the presence of multiple drug resistance genes compromises the therapeutic success by calculated antibiotic therapy. Various molecular and mass spectrometry-based approaches have been introduced in the recent past to speed up the provision of reliable resistance data. PCR- and sequencing-based approaches, though the most expensive, are frequently used for the detection of resistance genes even from clinical samples, bypassing the need for pure growth. Next-generation sequencing, based either on assessment of allelic single nucleotide polymorphisms or on the detection of non-ubiquitous resistance mechanisms, might allow for sequence-based bacterial resistance testing comparable to viral resistance testing in the long term. MALDI-TOF-MS, alone or in combination with molecular techniques, like PCR or minisequencing, has been found to provide the fastest resistance results from pure colonies or clinical samples. Nevertheless, the prohibitively high cost of these highly sophisticated diagnostic solutions limit their availability in resource-limited countries where multidrug resistance is on the rise (9). Molecular techniques like fluorescence in situ hybridization (FISH), which are more affordable, might be an option for resource-limited settings (10) until MALDI-TOF MS or sequence-based approaches become more affordable and easy to apply. However, all these technologies are far from being used in routine clinical microbiology laboratories. Also questionable is the repeatability of these tests in large clinical trials and studies, highlighting the need for standardization and creating quality assessment protocols for molecular tests. A network of diagnostic laboratories that can examine and evaluate the performance of molecular tests alongside the routine diagnostic tests is essential and should be the next step.
New treatment options
There is currently global alarm that the discovery void since 1990 has resulted in an empty antibiotic pipeline and the analogue development has also dried up. There were two fundamental mistakes in the way the scientific community looked at antibiotics, because of which we find ourselves where we are today. Firstly, there was an impression that infectious diseases were on the verge of elimination and, as a result, academic bacteriology distanced itself from antibiotic discovery, resulting in an overdependence on development of analogues. In the absence of any new molecules we have also exhausted the analogues that can be created out of existing products. Secondly, with the advent of genomics and proteomics, scientists believed that the next group of antibiotics would come out of the “OMICS”, which has not happened. Consequently, we have landed up with an inadequate pipeline of antibiotic molecules or their analogues. The last few decades did see development of new antibiotic molecules against Gram-positives, such as lineziolid and daptomycin, but no new molecules which are active against gram negatives have been reported and the analogues of Beta-lactamase inhibitors are providing hope to counter the emerging new resistant profiles among the Gram-negatives. Some Gram-negative bacteria, like NDM-1, producing K. pneumoniae and Acinetobacter baumanii, are showing resistance to most of the antibiotics, including carbapenems (11).
Unlike what was previously desired, current antibiotic discovery is focused on developing a potent, non-toxic antibiotic, having a narrow spectrum which is organism-specific. The challenge for the development of new antibiotics is to make the best use of the available technologies to ensure that the impact of emergence of resistance is fully factored into the development of next-generation antibacterial drugs and that drugs don’t fail owing to microbiological reasons which are now well known. Countries, like India, with high burden of Gram-negative infections in their healthcare settings, need one or more effective oral options for the treatment of ESBL Enterobacteriaceae and effective parenteral options for the treatment of carbapenemase (especially NDM-1) producing Enterobacteriaceae. While new classes of antibiotics are being developed to fight such infections, it is necessary to explore strategies to provide a bridge to new drugs, in particular by revisiting the use of existing (old) antibiotics, as well as their use in novel combination, either with another existing antibiotic or an approved drug. Polymyxins are used against XDR Gram-negative bacteria and while promising new drugs are currently in the pipeline, polymyxins should be explored for ways to increase the dosage for greater efficacy, while decreasing toxicity either through new formulations (for example, liposomal) or combinations. Colistin, which was discarded previously, is being increasingly used to treat carbapenem-resistant infections (12). Similarly, many groups are arguing repurposing the use of fosfomycin to treat drug-resistant N gonorrhea. In order that the old drugs are repurposed it is important to carry out in-vitro culture susceptibilities of resistant bugs to forgotten or re-purposed. It is also critical to understand the role of EDTA adjuvants in overcoming resistance in clinical practice.
Antibiotic combinations are used in clinical practice, but often without sufficient evidence to ensure clear clinical benefits. Given the high levels of resistance to carbapenems, randomized, controlled trials to decide the role of monotherapy versus combination therapy for carbapenem-resistant Gram-negative bacteria in order to preserve carbapenems and extend their life are the need of the hour. These combinations, though routinely used, need to be validated in controlled clinical trial settings (12).
Efforts are being made to identify antibiotic combinations that could restore activity of a given antibiotic that has lost, due to resistance or potentiate, its activity (the concept of ‘antibiotic resistance breaker’). By engaging in a systematic exploration of antibiotic combinations to identify synergistic additive, or even antagonist effects, it is highly plausible to unravel potent new combinations capable of overcoming certain resistance patterns. Whereas in vitro high throughput methods can readily be set up to identify such combinations, it has to be complemented with thorough PK/PD and in vivo experiments to validate their possible benefit over existing regimens. Revisiting PK/PD of old antibiotics, as many of them have been developed and registered with scarce and incomplete PK/PD analysis as compared to the current standards, has been listed as a priority area by organizations such as DNDi. This can lead to a reformulation or new regimens of such old drugs, with the aim to obtain the best efficacy while minimizing the emergence of resistance due to underdosage or inappropriate regimens. Such data can guide the development of alternative/improved formulations or regimens if they provide a clear therapeutic advantage over the old formulations or regimens. The HTS combinatorial screening and the PK/PD-in vivo platforms proposed can complement each other.
Although the search for new antibiotics continues, nanotechnology has revolutionized treatment of drug-resistant infections using the existing drugs. Nanocarriers have been used as a means to decrease resistance development by modifying delivery of vancomycin to treat VRSA. Carboxymethyl chitosan conjugated folic acid nanoparticles carrying vancomycin were shown to decrease the tolerance levels of resistant strains, acting as strong bactericid. Folic acid appeared to be the crucial part of the newly developed nanodelivery system (13). NS is becoming particularly interesting in this era of increased antibiotic-resistant and hospital-acquired bacterial infections. In addition, bacterial resistance to elemental silver is extremely rare. Nanosilver is the most commercialized nanomaterial with antibacterial properties, and its antibacterial effect is attributed to its mitochondrial toxicity due to interactions with thiol groups of internal membrane proteins causing oxidative stress. One of the pipelines in the development of successful antimicrobial therapy focuses on use of specific antimicrobial peptides. Local delivery of newly developed therapeutic agents which would not permit development of bacterial resistance represents an attractive option in improved antimicrobial therapy. It seems that antimicrobial peptides, associated with appropriate delivery system of nanosize range, could serve this purpose (14).
Strengthening antimicrobial stewardship within hospitals and beyond
Due to the shortage of new antibiotics being developed, it is has become imperative that current treatments are used judiciously by both healthcare professionals and members of the society/community. Antimicrobial stewardship programmes (AMSPs), which are multi-disciplinary, have been demonstrated to optimize antibiotic usage and reduce associated adverse events. Implementation of strong antimicrobial stewardship programmes has been helpful in checking the irrational use of antibiotics in many countries which have implemented AMSPs with varying degrees of success. Most countries found the antimicrobial stewardship leveling off after a certain period of time, thereby highlighting a need for research to identify factors that could help to sustain such programmes (15). Due to the complexity and heterogeneity of healthcare delivery across countries, the practice of antimicrobial stewardship is variable in LMICs. Even where policies were implemented and were found successful, full evaluation of cost-effectiveness, patient outcomes and impact of regulatory framework, etc., was not conducted. The ability to assess the effectiveness of an AMSP is often limited by external factors like drug shortages, outbreaks, changes in personnel and expansion of facilities or services. Presence of even one of these factors can mask the true intervention effect. It is also important to recognize that many AMSPs do not have data management or statistical support for programmatic evaluation. Thus, the lack of these resources can be a major barrier to conducting high-quality evaluations of AMSPs. There is therefore a strong case to support continued research into the impact of antibiotic stewardship programmes on process outcomes, to improve data recording and management systems and encourage increased research on the impact of such programmes on patient-centered outcomes (16).
Sociobehavioural research around antimicrobial resistance
Influencing human behavior in relation to antibiotics use is a complex process which includes taking into account factors such as knowledge, attitudes, peer pressure, and socioeconomic and sociobehavioral factors. Further, key messages and concepts are perceived in different ways by different individuals. While designing and implementing projects or programmes for behavioral change, with respect to antibiotics for professionals or consumers, it is helpful to consider that people in different stages of change are susceptible to different behaviour modification strategies. Understanding the linkage between specific economic conditions, social norms and attitudes relating to antibiotics is required in order to develop context-appropriate strategies. This requires interdisciplinary research linking clinical bacteriology, patient data and social science to investigate the factors influencing uptake and use of new treatments, how they are being implemented and how to ensure appropriate stewardship to prevent misuse and thus development of resistance. Application of social marketing principles needs to be tested in different country settings for its usefulness to bring down antibiotic use. Linking surveillance data to AMSP practices would be helpful in evaluating the impact of interventions to control/prevent the spread of resistant bacteria and at the same time assess the wider costs and benefits of behavior change strategies. Efforts need to be directed to develop and evaluate economic and regulatory models that balance the appropriate use of drugs and the mitigation of antimicrobial resistance, while appropriately rewarding innovation. This would require acquiring data from a range of sources – surveillance, healthcare, research and impact measures.
Making antibiotic resistance-related research attractive
To enable discovery of new diagnostics and next-generation novel antibiotics, a conducive investment climate and new business models that focus on sharing of the research costs for the development of new drugs and diagnostics among governments, the pharmaceutical industry, health systems, development agencies and large charitable foundations are urgently needed. Recently, Nesta Foundation along with Innovate UK has announced a £10 million prize to develop accurate, rapid, affordable, easy-to-use and readily available point of care diagnostic tests that will revolutionize antibiotic prescriptions. In India, Biotechnology Industry Research Assistance Council (BIRAC) has announced collaborations with Nesta, the UK’s Innovation Charity, to create an India-focused seed fund within the Discovery Award Programme of the Longitude Prize at Nesta, which would create a pipeline of Indian innovators for competing in the Longitude Prize.
In the field of drug development, there are a few industry-academia partnerships which are underway to improve the efficacy of existing drugs, but the capacity to discover new antibiotics within a short timeframe is severely lacking in both industry and academia. There is a need to develop a new sustainable global model for discovery and development of new antibiotics which is also a partnership between private and public sector. Drugs for Neglected Diseases Initiative (DNDI) has recently partnered with WHO to float a Global Antibiotic Research and Development Facility (GARD) which will expedite development of new antibiotics by catching low-hanging fruits, including repurposing old antibiotics by strengthening them with robust PK/PD data, reformulating the existing suboptimal formulations, combination trials with antibiotics resistance breakers and other adjuvants. Antibiotic research fund to fund Biotech/Pharma companies that work in the new-mechanism area and simplified regulations to allow for faster approvals for trials and extended patent rights would go a long way in getting the pharmaceutical industry interested in antibiotic discovery. The Generating Antibiotic Incentives Now (GAIN) Act was introduced in the USA and allows prolonged exclusivity and other provisions. This act could increase the value of antibiotics to companies, thereby encouraging companies to enter the field.
Efforts to make antibiotic related research and development attractive have to go in tandem with initiatives to create strong regulations and national preservation plans, to prevent misuse of new antibiotics when they become available. Kieny has suggested creating an intergovernmental consortium as a multipronged effort which promotes innovation, access and also controlled use. The proposed intergovernmental consortium will enable all the interested partners to contribute their skill as an innovator or manufacturer or consumer such that a framework for smooth transition of innovative products to the society is created (18).
Conclusions
“The best time to start was yesterday and next best is now.” To prevent regression to the pre-antimicrobial state of nature, antimicrobial resistance must be recognized as a significant problem and efforts undertaken to halt its growth. Understanding of antimicrobial resistance, both horizontally and vertically, underpins the development of strategies to contain antibiotic resistance and for assessment of the impact of interventions. The challenges posed by antimicrobial resistance are huge and gaps in our knowledge continue to pose a serious challenge in achieving the desired outcomes. Strong research efforts that lead to innovative solutions need to be supported at multiple levels. Significant progress has been made in this direction as discussed above. The challenge for the field now is to make the best use of the available technologies, information and expertise to ensure the impact of resistance is fully accounted for in the urgent development of much-needed interventions and solutions that are affordable and sustainable and can be adapted to different settings.
Biographies
Dr Soumya Swaminathan, Secretary, Department of Health Research (Ministry of Health & Family Welfare), Government of India, and Director General, ICMR, has vast experience in health research and research administration. Dr. Swaminathan was Director of the National Institute for Research in Tuberculosis (NIRT) in Chennai from 2012 to 2015. Following her MBBS from AFMC, Pune and MD in Pediatrics from AIIMS, New Delhi, further training included a Fellowship in Neonatology and Pediatric Pulmonology at the Children’s Hospital of Los Angeles, University of Southern California, USA and a Research Fellowship in the Dept. of Pediatric Respiratory Diseases, University of Leicester, UK. She joined the Tuberculosis Research Centre, Chennai in 1992 and has spent the past 24 years in health research. She was two years as Coordinator, Neglected Tropical Diseases at TDR.
Article available as PDF, click here
References
1. Davies J, Davies D. Origins and Evolution of Antibiotic Resistance. Microbiol. Mol. Biol. Rev. 2010;74:3417-433
2. Walia K. Emerging Problem of Antimicrobial Resistance in Developing Countries: Intertwining Socioeconomic Issues. Regional Health Forum 2003;7: 1-10
3. Review on Antimicrobial Resistance http://amr-review.org/
4. Okeke IN, Laxminarayan R, Bhutta ZA, Dus AG, Jenkins P, O’Brien TF, Pablos-Mendez A, Klugman KP. Antimicrobial resistance in developing countries. Part I: Recent trends and current status. Lancet Infect. Dis. 2005;5:481–493.
5. Aminov RI. A brief history of the antibiotic era: Lessons learned and challenges for the future. Front Microbiol. 2010;1:1-7
6. Global Antimicrobial Resistance Surveillance System (GLASS) http://www.who.int/drugresistance/surveillance/en/
7. Blair JMA, Webber MA, Baylay AJ, Ogbolu DO, Piddock LJV. Molecular mechanisms of antibiotic resistance. Nature Reviews Microbiology 2015;13:42–51
8. Loens K, Van Heirstraeten L, Malhotra-Kumar S, Goossens H, Ieven M. Optimal sampling sites and methods for detection of pathogens possibly causing community-acquired lower respiratory tract infections. J Clin Microbiol 2009;47:21–31.
9. Mishra RPN, Oviedo-Orta E, Prachi P, Rappuoli R, Bagnoli F. Vaccines and antibiotic resistance. Curr Opin Microbiol 2012;15:596–602.
10. Goodman C, Gould C, Karnes E, et al. The value of diagnostics innovation, adoption and diffusion into health care. The Lewin Group Reports. July 2005. http://www.lewin.com/~/media/Lewin/Site_Sections/Publications/ValueofDiagnostics.pdf (accessed Nov 1, 2013).
11. Laxminarayan R, et al. Antibiotic resistance—the need for global solutions. The Lancet Infectious Diseases. 2013;13:1057–98.
12. Biswas S, Brunel JM, Dubus JC, Reynaud-Gaubert M, Rolain JM. Colistin: an update on the antibiotic of the 21st century. Expert Rev Anti Infect Ther. 2012;10(8):917-34.
13. Chakraborty SP, Sahu SK, Mahapatra SK, et al. Nanoconjugatedvancomycin: new opportunities for the development of anti-VRSA agents. Nanotechnology 2010;21:1-9
14. Basnet P, Basnet NS. Nanodelivery Systems for Improved Topical Antimicrobial Therapy. Current Pharmaceutical Design 2013;19:7237-7243
15. Huttner B, Harbarth S, Nathwani D and ESGAP group. Success stories of implementation of antimicrobial stewardship: a narrative review. Clin Microbiol Infect 2014;20:954-962
16. MacDougall C, Polk RE. Antimicrobial Stewardship Programs in Health Care Systems Clin Microbiol Rev 2005;18:638–656
17. Lundborg CS, Tamhankar AJ. Understanding and changing human behaviour—antibiotic mainstreaming as an approach to facilitate modification of provider and consumer behavior. Upsala Journal of Medical Sciences. 2014;119:125–133
18. Kieny MP. Creating an intergovernmental consortium for new antibiotics: A new development model. “AMR Control 2015” http://www.globalhealthdynamics.co.uk/wp-content/uploads/2015/05/04_Kieny.pdf